 |
 |
- Search
| Intest Res > Volume 16(2); 2018 > Article |
|
Abstract
Background/Aims
Tens of trillions of microorganisms constitute the gut microbiota of the human body. The microbiota plays a critical role in maintaining host immunity and metabolism. Analyses of the gut microbial composition in Korea are limited to a few studies consisting of small sample sizes. To investigate the gut microbial community in a large sample of healthy Koreans, we analyzed the 16S ribosomal RNA of 4 representative bacterial genera Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium.
Methods
A total of 378 DNA samples extracted from 164 infants and 214 adults were analyzed using quantitative real-time polymerase chain reaction.
Results
Analysis of 16S ribosomal RNA of 4 representative bacterial genera Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium showed that the gut microbiota in infants had higher relative abundances of Bifidobacterium and Lactobacillus than that in adults, which was dominated by Bacteroides and Clostridium.
Conclusions
To the best of our knowledge, this was the first study evaluating the distinct characteristics of the microbial community of Korean infants and adults. The differences between the 2 populations suggest that external factors such as age, diet, and the environment are important contributing factors to the change in gut microbial composition during development.
Trillions of microorganisms colonize various regions of the human body, including the skin, mucosal surfaces, genitals, and gastrointestinal (GI) tract, outnumbering the cell numbers of their human hosts by 10 times.1 Immediately after delivery, newborn infants are exposed to a wide variety of microbes, and their gut microbiota is usually initially dominated by bifidobacteria, ultimately developing into a more diverse characteristic adult pattern.2,3 The composition and function of the gut microbiota rapidly develop to form a complex network that depends on the individual genome, nutrition, antibiotic use, and lifestyle, with variations among ethnicities and countries.4,5,6 The gut microbiota is associated with the regulation of metabolic pathways such as host-microbiota metabolism, signaling, and immune responses that connect the gut, liver, muscle, and brain.7 In addition, dysbiosis of microbial communities in both adults and infants has been shown to be linked to several functional gut disorders including obesity, IBD, type 1 diabetes, autism, necrotizing enterocolitis, and gastric and colon cancers.8,9,10,11,12,13 Several lines of evidence have shown that an abnormal increase or decrease in certain bacteria is associated with health and disease in humans.14,15,16,17,18 Based on these reports, 4 putative genera, Bifidobacterium, Lactobacillus, Bacteroides, and Clostridium, were chosen for analysis. As the gut microbiota plays an essential role in influencing the health of the host, it is important to first understand what constitutes a “healthy” gut microbiome. However, the majority of studies in the literature focusing on the characterization and assembly of the gut microbiota in healthy people have been limited to Western populations, with no extensive investigation of the microbial communities of the Korean population conducted to date.
Therefore, in the present study, we have focused on describing the large-scale profiles that constitute a healthy gut microbiota in Koreans to provide a detailed reference. Previous studies have indicated that the gut microbiota community differs before and after the first year of life.3 Furthermore, the gut microbial composition during infancy can have long-lasting effects on an individual's future health.19,20,21 The primary aim of the present study was to characterize the gut microbiome composition of healthy Koreans, using cultivation-independent molecular techniques. Fecal samples are often used to study the intestinal microflora because they are easily collected noninvasively. Previous studies have evaluated standard microbial cultures using selective media, but these methods lack reproducibility and most bacterial species cannot be cultured. However, progress in bacterial taxonomy based on molecular methods, especially the comparison of the 16S ribosomal RNA (16S rRNA) gene, has made culture-independent investigation the standard approach for identifying and phylogenetically classifying microbial communities.22,23
Although PCR amplification of the 16S rRNA gene in combination with pyrosequencing appears to be a useful approach for studying the microbial composition in an ecological environment such as the human gut, it is limited to providing only an estimation of the relative abundance without information on absolute bacterial density.24 In contrast, quantitative real-time PCR (qPCR) has the advantage of providing more accurate and quantitative information; however, it cannot provide any information on the relative abundance of the total bacterial composition.
Using this approach, we analyzed the gut microbiota of the 2 age groups with respect to the representative genera Lactobacillus, Bifidobacteria, Bacteroides, and Clostridium. The profiles established herein will provide a critical baseline for future studies investigating the Korean gut microbiome composition between infants and adults.
Fecal samples were collected from and donated by 378 healthy Korean individuals (164 infants and 214 adults) recruited at the Samsung Medical Center (Seoul, Korea). Clinical characteristics of the participants are shown in Table 1. The mean age of infants was 0.42±0.25 years, including 85 males and 79 females. The mean age of adults was 32.97±14.10 years, including 94 males and 120 females. Subjects who were overweight or had GI symptoms, a history of GI surgery, asthma, cancer, CD, IBS, IBD, diabetes, and neurological diseases were excluded. In addition, subjects with a history of antibiotic use or intentional probiotic intake were excluded in this study. Written informed consent was obtained from all subjects according to the institutional guidelines, and the protocol was approved by the Institutional Review Board of the participating hospitals (IRB number: 2013-12-061). All methods were performed in accordance with the relevant guidelines and regulations.
Test kits containing a sterile plastic tube (Sarstedt, Nümbrecht, Germany) with a screw cap to collect the stool samples were provided to the participants. The samples were collected by the participants themselves and immediately stored in home freezers until delivery to the experimental laboratory. The samples were stored at −80℃ until DNA was extracted.
Approximately 200 mg of each stool sample was fully homogenized with 0.1-mm diameter glass beads (Biospec, Orlando, FL, USA). Fecal DNA was subsequently extracted using commercial QIAamp stool DNA extraction kits (Qiagen, Valencia, CA, USA) according to the manufacturer's instructions. The concentration and quality of the extracted genomic DNA was determined by agarose gel electrophoresis and spectrophotometric analysis (NanoDrop Technologies, Wilmington, DE, USA). All extracted DNA samples were stored at −20℃ until use in further experiments.
To quantify the total bacteria and the bacteria of interest in the stool samples, qPCR was carried out using the ABI SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA) with primers amplifying the genes encoding 16S rRNA from specific bacterial groups (Table 2). qPCR for the 4 major genera representative of the gut microbiota (Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium) was performed as described previously25,26,27 to ensure broad bacterial coverage. Each qPCR was carried out in duplicate 20 µL reactions with 1 pM forward and reverse primers, 10 ng purified fecal DNA, and 1X SYBR qRT-PCR Master Mix (Takara Bio Inc., Shiga, Japan). The qPCR conditions were as follows: 4 minutes at 95℃, followed by 40 cycles of 95℃ for 15 seconds, 15 seconds at the appropriate annealing temperature, and 72℃ for 33 seconds. All qPCR plates included a “no-template” negative control for each primer set. All microbiology analyses of fecal samples were performed blindly, without knowledge of the subjects' clinical data. The concentrations of each bacteria in this study were normalized to 16S rRNA expression, the internal reference, in fecal samples using the comparative threshold cycle (CT) method.28 The concentrations of each genus in both groups were calculated as the quantitative abundance of bacterial ΔCT values:
To quantify each bacterial group, DNA from the respective reference strain was used for construction of a standard curve. The bacterial strains, efficiency, and coefficient of determination (R2 value) of the standard curve for qPCR are listed in Table 3. Culture conditions used to grow the standard bacterial strains are described in Supplementary Table 1. qPCR was performed as described above with serially diluted bacterial genomic DNA extracted from the reference strains using the QIAamp genomic DNA extraction kit (Qiagen), and the respective copy numbers of 16S rRNA were calculated. Each standard curve was constructed by plotting CT values against bacterial quantity (in 16S rRNA).
Clinical characteristics of subjects were expressed as the means±SD. Statistical analysis was performed using Graph-Pad Prism4 (GraphPad Software, San Diego, CA, USA). Abundance in the bacterial species between infants and adults was compared using the Mann-Whitney U test or the Kruskal-Wallis t-test, as appropriate. P<0.05 was regarded as statistically significant.
Fecal samples from 378 healthy Korean subjects were collected for analysis. Table 1 shows the characteristics of the subjects classified into 2 age groups (infants and adults). Statistically significant differences in age, height, weight, and BMI were observed between the 2 groups.
The study participants consisted of 378 healthy Koreans, including 164 infants and 214 adults. In order to analyze the fecal microbiota of this population, we targeted the 4 representative genera of the human gut microbiota to ensure coverage of fecal bacteria composition. Fig. 1 shows the relative abundance of the fecal microbiota groups as determined by qPCR from all subjects enrolled in this study. The most abundant genus overall was Bacteroides, accounting for over 40% of the total microbiota, followed by Bifidobacterium and Clostridium with an average abundance of 28.69% and 25.37%, respectively (Fig. 1A). The proportion of Lactobacillus was below 3% in most of the samples. Lactobacillus was the only group with a significantly lower level (P<0.0001) of normalized copy number by qPCR among the 4 bacteria genera in the overall population sample of healthy Koreans (Fig. 1B).
To identify differences in the characteristics of the gut microbiota according to age, we compared the relative composition of each of the 4 genera in infants and adults. According to our predictions, the average proportion of Bifidobacterium was highest (52.46%) in gut microbiota in infants (Fig. 2A), which is in line with previous observations.29,30 The relative abundance of Bifidobacterium was significantly higher than that of the other genera in infants (Fig. 2B). Whereas Bacteroides was the most dominant (66.16%) genus in the gut microbiota of adults (Fig. 2C), which is also supported by previous studies.31,32 When we compared the difference of the gut microbiota composition according to sex, the levels of Lactobacillus and Bacteroides of adult females were significantly higher than those of males (data not shown). There were no significant differences in the composition of the other bacteria between males and females according to relative abundance and normalized values. In adults, the normalized values of Bacteroides and Lactobacillus were significantly higher and lower than those of other genera, respectively (Fig. 2D). Overall, these results indicated that Bifidobacterium is the initial colonizer of the gut microbiota, with a subsequent switch to Bacteroides as the dominant bacteria after the first year of life.
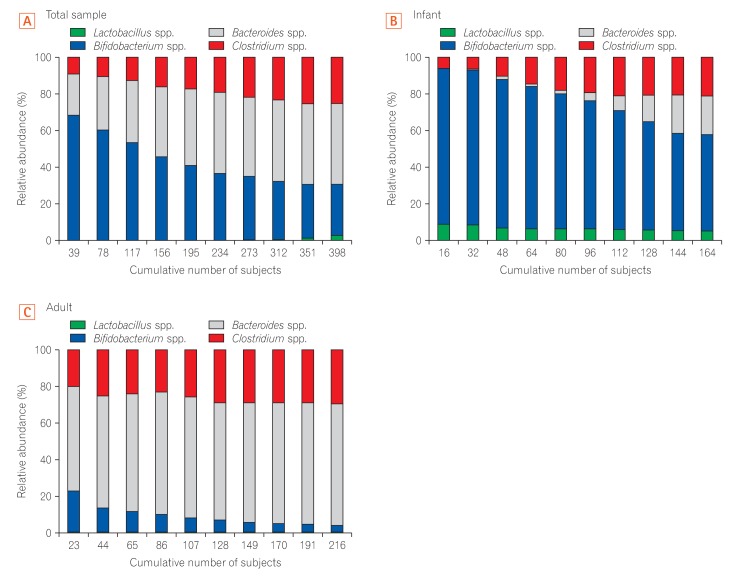
To characterize the composition pattern of the 4 genera according to age in the Korean population, we analyzed the cumulative change in the relative abundance of each genera in the gut of healthy Korean subjects. We classified all Korean subjects on the basis of accumulated values and compared the relative abundance of the gut microbiota for each accumulated point. Although Bifidobacterium was enriched up to a high accumulated value of 30%, Bacteroides gradually became dominant towards the lower accumulated value of the entire sample (Fig. 3A). In contrast, analysis of the gut microbiota composition in infants showed that Bifidobacterium was highly enriched at all accumulated values (Fig. 3B). The proportion of Lactobacillus was lowest in the total sample and in adults. In contrast, Bifidobacterium was low and the proportion of Bacteroides was high at all accumulated values in the gut microbiota community in adults (Fig. 3C). In all cumulative graphs, Clostridium became gradually more enriched towards the lower accumulated value.
During adult life, the community composition of the gut microbiota could be influenced by host genetics, health, diet, probiotics, and aging.8,33 Although there have been several reports on investigations of the gut microbiome in Western and in some Asian populations,8,32,34 little is known about the healthy Korean gut microbiota. To identify the gut microbiome composition of the healthy Korean population, in the present study, we analyzed the gut microbiota composition of 378 Koreans using a molecular approach. Indeed, several bacteria that had never been isolated from human feces were detected by PCR in the present study. Previous studies have analyzed the levels of Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium genera that commonly inhabit in the human colon in feces by PCR.14,35,36
According to previous clinical studies, the levels of certain bacteria may vary in health and disease.15,16,17,18,37,38
Lactobacillus is the predominant beneficial bacteria in adults. The level of Lactobacillus is generally lower in patients with IBS than in healthy controls.39
Bifidobacterium have been considered to be the most essential organisms for infants and also constitute one of the most important organisms in the colonic flora of healthy children and adults.3 In particular, the abundance of Bifidobacterium is decreased in patients with some functional GI disorders such as IBS and IBD.15,18 Some Lactobacillus and Bifidobacterium strains are used as probiotics, showing a beneficial effect in the prevention of inflammation in the intestine16 and the ability to eliminate harmful factors impeding the repair of damaged tissue. Hence, probiotics play a key role in aging, regulation of brain gene expression, and inflammation.17,40 Dethlefsen et al.33 found that a small number of genera within the Bacteroidetes and Firmicutes phyla accounted for the majority of gut microbial communities. Bacteroides and Clostridium belong to the phyla Bacteroidetes and Firmicutes, respectively and have been found to be increased in abundance in patients with IBS.37,38 These studies imply that changes in these 4 bacterial genera: Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium are closely related to human health and disease.
Based on this background, we examined the average distributions of Lactobacillus, Bifidobacterium, Bacteroides, and Clostridium in the gut microbiota of healthy Koreans. The most abundant genus in the total samples was Bacteroides, at 42.98%, followed by Bifidobacterium and Clostridium at 28.69% and 25.37%, respectively. Nam et al.41 also reported that Bacteroides was 1 of the 3 most abundant genera in the Korean gut microbiota. Moreover, Arumugam et al.42 found that levels of Bacteroides were relatively high in the fecal metagenomes of individuals from 4 countries, thus classifying Bacteroides as an enterotype.
The diversity of the genus Bifidobacterium in the infant gut microbiota is currently not well understood. In culture-based studies of the gut microbiota associated with aging, Mitsuoka3 showed that Bifidobacteria were the major organisms in the colonic flora of healthy children and adults. Similarly, the fecal samples of the infants in the present study showed a predominance of Bifidobacterium using a cultureindependent system. In agreement with our results, Turroni et al.43 also found a large proportion of Bifidobacterium in the gut microbiota of 11 infants. These studies reinforce the notion that Bifidobacteria are a predominant component of the infant gut microbiota. Bacterial populations within the infant gut are influenced by many factors such as mode of delivery, breastfeeding versus formula feeding, and the use of antibiotics.44,45 Infants tend to have an unstable, less diverse gut microbiota composition than adults. Following the consumption of solid food, the gut microbial community of the infant begins to develop towards the adult pattern, which differs from that of infants. Indeed, in our study, adults typically had a gut community dominated by Bacteroides . Surprisingly, we confirmed that the gut microbiota composition of our relatively small random sample showed a similar distribution of other bacteria including Bacteroides in this population. These findings strongly support the notion that healthy Koreans tend to show a common gut microbiota composition.
Taken together, we have demonstrated that the average relative abundance of 4 genera in the gut microbiota can be analyzed at a large scale in healthy Koreans using a culture-independent method from fecal samples. Although we could not evaluate the effects of dietary patterns and environmental factors on the microbiota patterns identified in the present study, we believe that these factors should be taken into consideration in future studies. As only a limited number of individuals have been subjected to GI microbiota analysis thus far, our data contribute to gaining a comprehensive understanding of the gut microbiota composition of healthy Koreans. These results will be beneficial to future evaluations of the causal relationship between gut microbiota and various diseases, such as IBD, obesity, colon cancer, asthma, and atopy in Korean patients, and should help to establish a reference for the gut microbiota composition in healthy Korean individuals.
References
1. Eckburg PB, Bik EM, Bernstein CN, et al. Diversity of the human intestinal microbial flora. Science 2005;308:1635-1638.PMID: 15831718.



2. Dominguez-Bello MG, Costello EK, Contreras M, et al. Delivery mode shapes the acquisition and structure of the initial microbiota across multiple body habitats in newborns. Proc Natl Acad Sci U S A 2010;107:11971-11975.PMID: 20566857.



3. Mitsuoka T. Bifidobacteria and their role in human health. J Ind Microbiol Biotechnol 1990;6:263-267.

4. Palmer C, Bik EM, DiGiulio DB, Relman DA, Brown PO. Development of the human infant intestinal microbiota. PLoS Biol 2007;5:e177. PMID: 10.1371/journal.pbio.0050177. PMID: 17594176.



5. Suzuki TA, Worobey M. Geographical variation of human gut microbial composition. Biol Lett 2014;10:20131037PMID: 10.1098/rsbl.2013.1037. PMID: 24522631.



6. David LA, Maurice CF, Carmody RN, et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 2014;505:559-563.PMID: 24336217.



7. Nicholson JK, Holmes E, Kinross J, et al. Host-gut microbiota metabolic interactions. Science 2012;336:1262-1267.PMID: 22674330.


8. Turnbaugh PJ, Hamady M, Yatsunenko T, et al. A core gut microbiome in obese and lean twins. Nature 2009;457:480-484.PMID: 19043404.


9. de la Cochetiere MF, Piloquet H, des Robert C, Darmaun D, Galmiche JP, Roze JC. Early intestinal bacterial colonization and necrotizing enterocolitis in premature infants: the putative role of Clostridium. Pediatr Res 2004;56:366-370.PMID: 15201403.


10. Fell JM. Neonatal inflammatory intestinal diseases: necrotising enterocolitis and allergic colitis. Early Hum Dev 2005;81:117-122.PMID: 15707723.


11. Finegold SM. Therapy and epidemiology of autism: clostridial spores as key elements. Med Hypotheses 2008;70:508-511.PMID: 17904761.


12. O'Keefe SJ. Nutrition and colonic health: the critical role of the microbiota. Curr Opin Gastroenterol 2008;24:51-58.PMID: 18043233.


13. Marchesi JR, Holmes E, Khan F, et al. Rapid and noninvasive metabonomic characterization of inflammatory bowel disease. J Proteome Res 2007;6:546-551.PMID: 17269711.


14. Penders J, Vink C, Driessen C, London N, Thijs C, Stobberingh EE. Quantification of Bifidobacterium spp., Escherichia coli and Clostridium difficile in faecal samples of breast-fed and formula-fed infants by real-time PCR. FEMS Microbiol Lett 2005;243:141-147.PMID: 15668012.



15. Giaffer MH, Holdsworth CD, Duerden BI. The assessment of faecal flora in patients with inflammatory bowel disease by a simplified bacteriological technique. J Med Microbiol 1991;35:238-243.PMID: 1941994.


16. Madsen KL, Doyle JS, Jewell LD, Tavernini MM, Fedorak RN. Lactobacillus species prevents colitis in interleukin 10 gene-deficient mice. Gastroenterology 1999;116:1107-1114.PMID: 10220502.


17. Mencarelli A, Distrutti E, Renga B, et al. Probiotics modulate intestinal expression of nuclear receptor and provide counter-regulatory signals to inflammation-driven adipose tissue activation. PLoS One 2011;6:e22978. PMID: 10.1371/journal.pone.0022978. PMID: 21829567.



18. Kerckhoffs AP, Samsom M, van der Rest ME, et al. Lower Bifidobacteria counts in both duodenal mucosa-associated and fecal microbiota in irritable bowel syndrome patients. World J Gastroenterol 2009;15:2887-2892.PMID: 19533811.



19. Cox LM, Yamanishi S, Sohn J, et al. Altering the intestinal microbiota during a critical developmental window has lasting metabolic consequences. Cell 2014;158:705-721.PMID: 25126780.



20. Kostic AD, Gevers D, Siljander H, et al. The dynamics of the human infant gut microbiome in development and in progression toward type 1 diabetes. Cell Host Microbe 2015;17:260-273.PMID: 25662751.



21. Suau A, Bonnet R, Sutren M, et al. Direct analysis of genes encoding 16S rRNA from complex communities reveals many novel molecular species within the human gut. Appl Environ Microbiol 1999;65:4799-4807.PMID: 10543789.



22. Kim SE, Choi SC, Park KS, et al. Change of fecal flora and effectiveness of the short-term VSL#3 probiotic treatment in patients with functional constipation. J Neurogastroenterol Motil 2015;21:111-120.PMID: 25537674.




23. Kwok LY, Zhang J, Guo Z, et al. Characterization of fecal microbiota across seven Chinese ethnic groups by quantitative polymerase chain reaction. PLoS One 2014;9:e93631. PMID: 10.1371/journal.pone.0093631. PMID: 24699404.



24. Carroll IM, Chang YH, Park J, Sartor RB, Ringel Y. Luminal and mucosal-associated intestinal microbiota in patients with diarrhea-predominant irritable bowel syndrome. Gut Pathog 2010;2:19PMID: 10.1186/1757-4749-2-19. PMID: 21143915.



25. Wang Y, Ametaj BN, Ambrose DJ, Gänzle MG. Characterisation of the bacterial microbiota of the vagina of dairy cows and isolation of pediocin-producing Pediococcus acidilactici. BMC Microbiol 2013;13:19PMID: 10.1186/1471-2180-13-19. PMID: 23356904.



26. Matsuki T, Watanabe K, Fujimoto J, et al. Development of 16S rRNA-gene-targeted group-specific primers for the detection and identification of predominant bacteria in human feces. Appl Environ Microbiol 2002;68:5445-5451.PMID: 12406736.



27. Rinttilä T, Kassinen A, Malinen E, Krogius L, Palva A. Development of an extensive set of 16S rDNA-targeted primers for quantification of pathogenic and indigenous bacteria in faecal samples by real-time PCR. J Appl Microbiol 2004;97:1166-1177.PMID: 15546407.


28. Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 2008;3:1101-1108.PMID: 18546601.



29. Sjögren YM, Jenmalm MC, Böttcher MF, Björkstén B, Sverremark-Ekström E. Altered early infant gut microbiota in children developing allergy up to 5 years of age. Clin Exp Allergy 2009;39:518-526.PMID: 19220322.


30. Hopkins MJ, Sharp R, Macfarlane GT. Age and disease related changes in intestinal bacterial populations assessed by cell culture, 16S rRNA abundance, and community cellular fatty acid profiles. Gut 2001;48:198-205.PMID: 11156640.



31. Tap J, Mondot S, Levenez F, et al. Towards the human intestinal microbiota phylogenetic core. Environ Microbiol 2009;11:2574-2584.PMID: 19601958.


32. Kurokawa K, Itoh T, Kuwahara T, et al. Comparative metagenomics revealed commonly enriched gene sets in human gut microbiomes. DNA Res 2007;14:169-181.PMID: 17916580.




33. Dethlefsen L, Huse S, Sogin ML, Relman DA. The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biol 2008;6:e280. PMID: 10.1371/journal.pbio.0060280. PMID: 19018661.



34. Gill SR, Pop M, Deboy RT, et al. Metagenomic analysis of the human distal gut microbiome. Science 2006;312:1355-1359.PMID: 16741115.



35. Bélanger SD, Boissinot M, Clairoux N, Picard FJ, Bergeron MG. Rapid detection of Clostridium difficile in feces by real-time PCR. J Clin Microbiol 2003;41:730-734.PMID: 12574274.



36. Huijsdens XW, Linskens RK, Mak M, Meuwissen SG, Vandenbroucke-Grauls CM, Savelkoul PH. Quantification of bacteria adherent to gastrointestinal mucosa by real-time PCR. J Clin Microbiol 2002;40:4423-4427.PMID: 12454130.



37. Swidsinski A, Weber J, Loening-Baucke V, Hale LP, Lochs H. Spatial organization and composition of the mucosal flora in patients with inflammatory bowel disease. J Clin Microbiol 2005;43:3380-3389.PMID: 16000463.



38. Kassinen A, Krogius-Kurikka L, Mäkivuokko H, et al. The fecal microbiota of irritable bowel syndrome patients differs significantly from that of healthy subjects. Gastroenterology 2007;133:24-33.PMID: 17631127.


39. Malinen E, Rinttilä T, Kajander K, et al. Analysis of the fecal microbiota of irritable bowel syndrome patients and healthy controls with real-time PCR. Am J Gastroenterol 2005;100:373-382.PMID: 15667495.


40. Distrutti E, O'Reilly JA, McDonald C, et al. Modulation of intestinal microbiota by the probiotic VSL#3 resets brain gene expression and ameliorates the age-related deficit in LTP. PLoS One 2014;9:e106503. PMID: 10.1371/journal.pone.0106503. PMID: 25202975.



41. Nam YD, Jung MJ, Roh SW, Kim MS, Bae JW. Comparative analysis of Korean human gut microbiota by barcoded pyrosequencing. PLoS One 2011;6:e22109. PMID: 10.1371/journal.pone.0022109. PMID: 21829445.



42. Arumugam M, Raes J, Pelletier E, et al. Enterotypes of the human gut microbiome. Nature 2011;473:174-180.PMID: 21508958.



43. Turroni F, Peano C, Pass DA, et al. Diversity of bifidobacteria within the infant gut microbiota. PLoS One 2012;7:e36957. PMID: 10.1371/journal.pone.0036957. PMID: 22606315.



44. Lozupone CA, Stombaugh J, Gonzalez A, et al. Meta-analyses of studies of the human microbiota. Genome Res 2013;23:1704-1714.PMID: 23861384.



45. Jakobsson HE, Jernberg C, Andersson AF, Sjölund-Karlsson M, Jansson JK, Engstrand L. Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS One 2010;5:e9836. PMID: 10.1371/journal.pone.0009836. PMID: 20352091.



Fig. 1
Pie chart of gut microbiota composition of the 4 genus levels examined in fecal samples of healthy Korean individuals. (A) Pie chart indicating changes in the average relative abundance of 4 bacterial genera in stool samples from 378 subjects. (B) Normalized values were calculated using the comparative threshold cycle (CT) method. The normalized value of Lactobacillus in fecal specimens from all subjects was significantly lower than those of other genera. aP<0.001.

Fig. 2
Pie chart of gut microbiota composition at the 4 genus levels examined in fecal samples of infants and adults. (A) Pie chart indicating differences in the relative abundance of 4 bacteria genera in stool samples from 164 infants. (B) The normalized value of Bifidobacterium in fecal samples of infants was significantly higher than those of other genera. (C) Pie chart indicating differences in the relative abundance of 4 bacteria genera in stool samples from 214 adults. (D) The normalized value of Bacteroides in fecal samples of adults was significantly higher than those of other genera. aP<0.001.
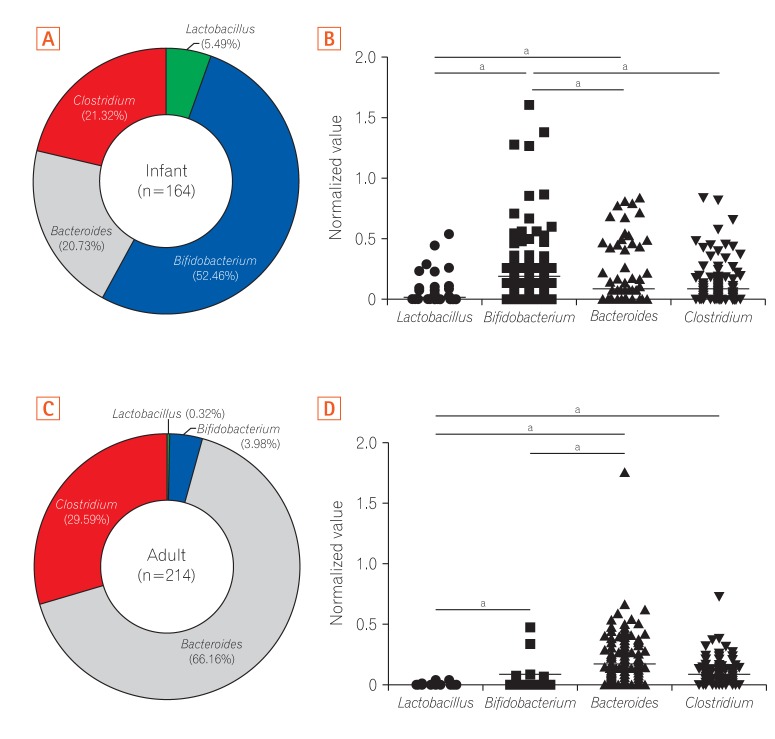
Fig. 3
Average cumulative abundance of the 4 genera in healthy Korean individuals. Distribution of 4 bacterial genera revealing the cumulative composition of microbial diversity in the 3 groups: total sample (A), infant (B), and adult (C). The X-axis of each graph represents the accumulated number of samples, divided by the total number of subjects and Y-axis of each graph indicates the relative abundance. The total sample includes both infants and adults.




 .
.

